 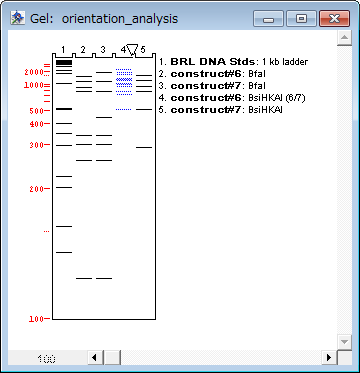
However, determination of the N-terminal amino acid sequence in the fusion protein shows that the polypeptide chain is heterogeneous and shortened by 4 and 5 amino acid residues. It is seen that Sarc-Ob is synthesized in the cells with a high yield and can be isolated on Ni 2+-NTA- agarose with a purity over 90% directly from the cell lysate in one step. Figure 1 B (lines 1 and 2) shows the level of synthesis and purification of the fusion protein sarcotoxin-obelin. Thus the formation of inclusion bodies prevents proteolytic degradation of the non-structured regions of the polypeptide chain. coli cells with the formation of inclusion bodies and that, after its isolation from these bodies, apoobelin, despite the absence of a rigid structure, can be transformed into the active form by incubation with the substrate. It was previously shown that apoobelin is synthesized in E. The obtained data show that endogenous proteases present in the cell lysate primarily cleave the polypeptide chain at the region connecting the functional parts of the fusion protein to the parts without a rigid structure, resulting from the character of the amino acid sequence (impossibility of folding).įor the fusion sarcotoxin-obelin we created a gene construct on the basis of photoprotein obelin. Glutation-S-transferase under thrombin treatment fully retains its activity and is resistant to thrombin action ( Fig. Sarcotoxin is a short polypeptide chain consisting of 41 amino acid residues without a rigid spatial structure and, as a result, easily undergoes proteolytical cleavage. Treatment of the isolated GST-Sarc and GST with thrombin results in complete cleavage of the fusion protein with the formation of GST and, apparently, in complete degradation of sarcotoxin. In contrast to 30 kDa polypeptide, the 35 kDa polypeptide from line 4 is copurified with 50 kDa major protein during Ni 2+-NTA chromatography (not shown)Īfter isolation of the fusion protein a further decrease of the amount of full-sized protein takes place in the column with glutation-Sepharose, while the amount of glutation-S-transferase increases ( Fig. Fluorescence of GFP module is associated with the major bands of 30 kDa on line 2 and of 35 kDa on line 4. Line 1, inclusion bodies of GFP-Ob I in 6 M urea 2, renatured GFP-Ob I 3, purified proteins from line 2 after Ni 2+-NTA chromatography 4, renatured from inclusion bodies GFP-Ob II. (Panel D) Green fluorescent protein-Obelins (GFP-Ob I and GFP-Ob II). Line 1, inclusion bodies 2, purified proteins after Ni 2+-NTA chromatography 3, proteins from line 2 purified after gel chromatography with Sephacryl S300 (Panel C) 6His-Dihydrofolate reductase-Obelin (DHFR-Ob). Line 1, inclusion bodies 2, purified protein after Ni 2+- NTA chromatography Line 1, 30,000 g supernatant after sonication 2, undigested, and 3, digested (with thrombin) proteins purified by chromatography on Glutathione Sepharose 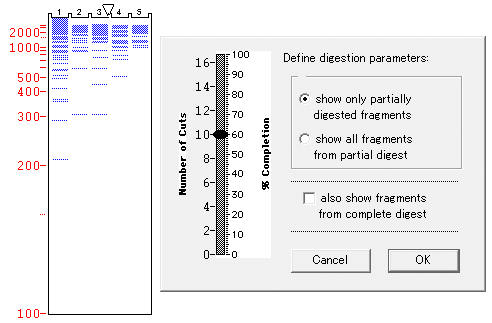
(Panel A) Glutathione-S-transferase - Sarcotoxin (GST-Sarc). Molecular mass standards 90, 66, 44, 30, 20 and 14 kDa are shown as “M”. coli fusion proteins by SDS-gradient polyacrylamide (10-20%) gel electrophoresis. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
